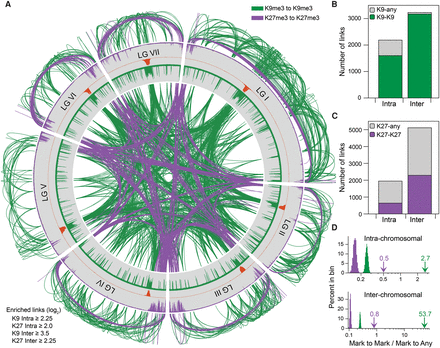
Predicted networks of H3K9me3-marked and H3K27me2/3-marked regions. (A) Plot of frequently observed contacts between heterochromatic regions from all seven chromosomes (LGI–LGVII) at 10-kb resolution. Green and purple links indicate frequent contacts between regions enriched with H3K9me3 or H3K27me2/3, respectively. Links inside and outside the plot show inter- and intra-chromosomal associations, respectively. The near absence of subtelomeric interactions between H3K27me2/3-enriched regions on LGV may reflect the incomplete assembly of the rDNA repeats on the LGV left arm. (B) Summary of observed levels of strong intra- and inter-chromosomal links between regions enriched with H3K9me3 (green bars) or between H3K9me3-enriched regions and regions without H3K9me3 (gray bars), generated with the script matrix2CircosK9Quant.py. (C) Summary of strong intra- and inter-chromosomal links between regions enriched with H3K27me2/3 (purple bars) or between regions enriched with H3K27me2/3 and regions without H3K27me2/3 (gray bars), generated with the script matrix2CircosK27Quant.py. (D) Distributions of ratios of interactions of like (Mark to Mark) or unlike (Mark to Any) regions from 10,000 randomly generated arrangements of strong intra- and inter-chromosomal contacts originating from H3K9me3-enriched regions (green) or H3K27me2/3-enriched regions (purple), produced by the scripts K9RandomizeInter.py, K9RandomizeIntra.py, K27RandomInter.py, and K27RandomizeIntra.py. The observed ratios of specific to nonspecific contacts for the H3K9me3 and H3K27me2/3 networks are denoted with green and purple arrows, respectively, and correspond to the plots in B and C.











