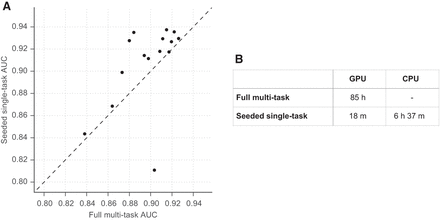
Basset leverages large-scale public data to inform additional data set learning. (A) The scatter plot shows AUC for 15 data sets achieved by the full model trained on all 164 cell types on the x-axis and AUC achieved by a procedure to simulate studying that data set alone on the y-axis. To study the data set alone, we pretrain a model on 149 cells (after removing these 15), seed training of the additional cell with that model's parameters, and perform a single training pass through the new data. This rapid procedure was effective for all but one data set (HRCEpiC, renal cortical epithelial cells), for which multitask training with the many other similar epithelial cells was beneficial. The AUC improvement for many cells suggests that our full model may benefit from increased capacity or decreased regularization. (B) The seeded training procedure is far faster on the GPU and allows for feasible CPU training.











