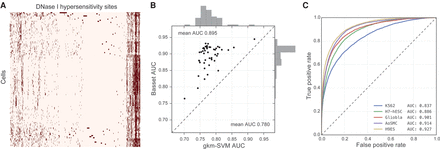
Basset accurately predicts cell-specific DNA accessibility. (A) The heat map displays hypersensitivity of 2 million DNase I hypersensitive sites (DHSs) mapped across 164 cell types. We performed average linkage hierarchical clustering using Euclidean distance to both cells and sites. (B) The scatter plot displays AUC for 50 randomly selected cell types achieved by Basset and the state-of-the-art approach gkm-SVM, which uses support vector machines. (C) The ROC curves display the Basset false-positive rate versus true-positive rate for five cells, selected to represent the 0.05, 0.33, 0.50, 0.67, and 0.95 quantiles of the AUC distribution.











