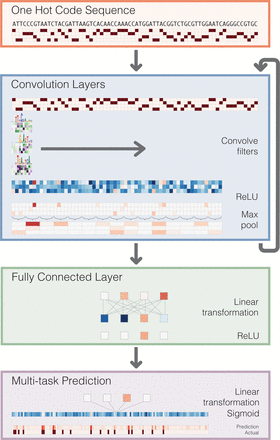
Deep convolutional neural network (CNN) for DNA sequence analysis. Basset predicts the cell-specific functional activity (here DNase I hypersensitivity) of sequences. First, we convert the sequence to a “one hot code” representation, where each position has a four-element vector with one nucleotide's bit set to one. Convolution layers proceed by scanning weight matrices across the input matrix to produce an output matrix with a row for every convolution filter and a column for every position in the input (minus the width of the filter). We apply a rectified linear unit (ReLU) nonlinear transformation to the convolution output and pool by taking the maximum across a window of adjacent positions. The first convolution layer operates directly on the one hot coding of the input sequence, making the convolution filters akin to the common bioinformatics tool position weight matrices. Subsequent convolution layers consider the orientations and spatial distances between patterns recognized in the previous layer. Fully connected layers perform a linear transformation of the input vector and apply a ReLU. The final layer performs a linear transformation to a vector of 164 elements that represents the target cells. A sigmoid nonlinearity maps this vector to the range zero to one, where the elements serve as probability predictions of DNase I hypersensitivity, to be compared via a loss function to the true hypersensitivity vector.











