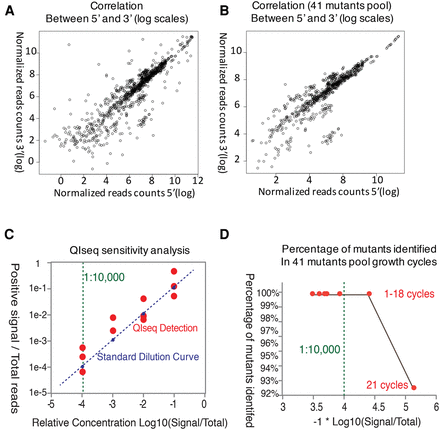
Reproducibility and sensitivity of QIseq. Insertion sites identified during sequencing of multiple single and pooled piggyBac lines were compared across biological replicates and time points. (A) Correlation plot of insertion sites identified by 5′ and 3′ libraries in small pool of 41 piggyBac insertions. (B) Correlation plot of the same insertion sites identified in biological replicates. (C) QIseq sensitivity analysis of the gDNA of PB-3 mutant line was diluted as 1:10, 1:100, 1:1000, 1:10,000 and mixed with three different mutant lines as biological reps. QIseq identifying and monitoring sites over a >10,000-fold dynamic range of sequence counts. (D) Percentage of mutants identified in small pool growth cycles, over a >10,000-fold dynamic range of sequence counts in 18 cycles.











