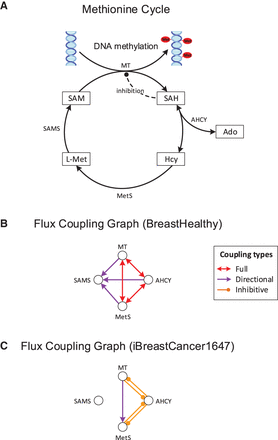
Control of methylation by the AHCY driver reaction. (A) Central reactions of the methionine cycle, essential for methylation of DNA, RNA, and proteins. AHCY is identified by our method as a metabolic switch for urothelial and breast cancer. It controls methylation by regulating the level of produced SAH, which inhibits methyltransferases. (B) Flux coupling graph of the reactions from A in healthy breast tissue. Full coupling between AHCY and methyltransferase ensures that SAH is produced and consumed at equal rates, avoiding its accumulation and DNA hypomethylation. For clarity, only the coupling relations between the reactions from A are shown. (C) Flux coupling graph of the reactions from A in the breast cancer network. The coupling relations differ from those of the healthy breast tissue network due to the differences in the included reactions and metabolites. AHCY is no longer fully coupled to methyltransferase and methionine synthase, allowing for independent fluxes of the reactions and varying SAH levels. Inhibitive coupling between AHCY and methyltransferase implies that elevated flux of either reaction inhibits flux of the other reaction due to accumulation of the common product SAH, leading to DNA hypomethylation and cancer, which is in agreement with experimental findings (Results). (Ado) Adenosine, (Hcy) Homocysteine, (L-Met) L-Methionine, (SAH) S-adenosylhomocysteine, (AHCY) adenosylhomocysteinase, (SAM) S-adenosylmethionine, (SAMS) SAM synthase, (MetS) methionine synthase, (MT) methyltransferase.











