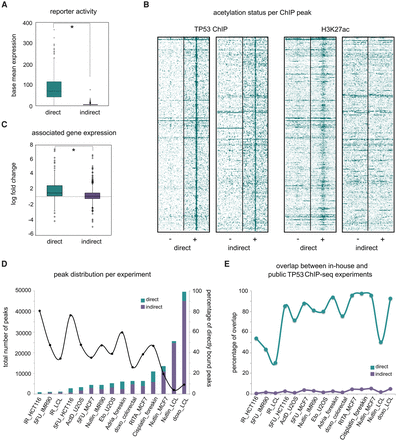
Only directly bound peaks behave as enhancers. (A) The average base mean expression values indicate that the direct and indirect peaks show significantly different reporter activity levels (P-value = 3.28 × 10−51). (B) Comparison of TP53 ChIP-seq signal and H3K27ac ChIP-seq signal between positives and negatives. Peaks are extended to 2000 bp each side. Heatmaps show the raw tag count coverage per peak. (C) The average differential expression of genes near (<20 kb) peaks: (*) P-value 4.73 × 10−8. (D) Comparison of the proportion of direct versus indirect ChIP-seq peaks across 15 publicly available TP53 ChIP-seq data sets. Experiments are ordered along the x-axis based on total number of peaks called. (E) Directly bound peaks agree, but indirectly bound do not, between in-house ChIP-seq peaks and other data sets. The percentage overlap is compared to the in-house peaks.











