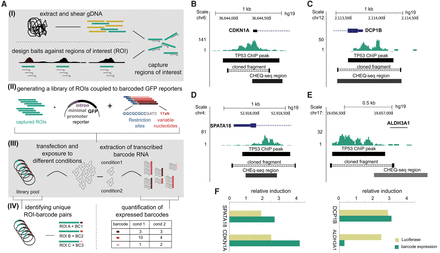
Overview of the CHEQ-seq reporter assay. (A) (I) Genomic DNA is sheared and custom baits are used to capture the regions of interest (ROI). (II) Captured ROIs are cloned into a reporter library consisting of a GFP-based reporter linked to a pool of 17 × 109 barcodes. (III) The reporter library is transfected under various conditions, after which the RNA of transcribed barcodes is extracted. (IV) Randomly coupled ROI-barcode couples are identified using PacBio sequencing, and barcode expression is measured using Illumina short-read sequencing. (B–E) Four TP53 ChIP-seq peaks comparing the CHEQ-seq barcode level with luciferase activity of a manually cloned fragment. (F) CHEQ-seq and luciferase induction only agree when they both overlap with the ChIP-seq peak summit.











