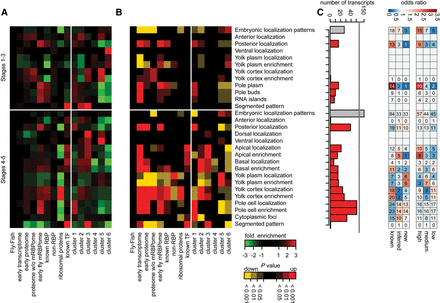
Localization of RBP-enocding mRNAs in early Drosophila embryos. RNA fluorescent in situ hybridization information from the Fly-FISH database (Lécuyer et al. 2007) was used to empirically calculate overrepresented transcript localization annotation terms by random sampling relative to the Fly-FISH database. Heat map indicating fold changes (A) and P values (B) for each subset for embryonic localization terms in embryonic developmental stages 1–3 and 4–5. Early transcriptome: protein coding genes expressed (FPKM > 0) in 0–2-h embryos; early proteome: proteins identified by whole 0–2-h embryo mass spectrometry; proteome without mRBPome: does not contain genes identified by mRBPome capture; known RBPs: RBPs selected by GO-term and RBD within the transcriptome excluding ribosomal proteins; non-RBP: transcriptome without known RBPs; ribosomal proteins: all ribosomal proteins; known TF: transcription factors from FlyTF.org database. Clusters as in Fig. 4A. Relative subset representation can be assessed in Supplemental Figure S5A. (C) The number of transcripts of the early fly mRBPome for each localization category. Gray bars = number for all transcripts with spatially restricted (≠ ubiquitous) embryonic localization terms within stages 1–3 and 4–5, respectively; red bars = number of transcripts in individual localization categories. Odds ratio of subcategories relative to the early fly mRBPome. Subcategories represent classification from Figures 2D and 3A.











