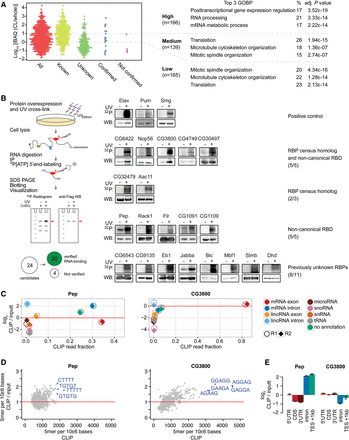
Validation of direct RNA interaction of candidate RBPs. (A) Protein enrichment (log10 iBAQ ratios of proteins in oligo[dT] precipitate versus whole-embryo proteins). Proteins are divided by enrichment score (high = >10-fold enrichment, middle = 1–10-fold enrichment, low = <1-fold enrichment) (six proteins [two validation candidates] missed whole embryo detection and could not be added here). The three most enriched GO-terms for biological processes (GOBP) for each category are shown on the right. Validation candidates were chosen throughout the enrichment ratio range, and were not annotated as RNA-binding in GOMF or having classical RBDs (except Pep). (B) Scheme describing the experimental approach. Epitope-tagged candidate RBPs were expressed in transiently or stably transfected Drosophila S2 cells. After 254-nm UV-crosslinking, cell lysis, and RNase digestion, the immunoprecipitated crosslinked protein-RNA complexes were radiolabeled by T4 PNK phosphorylation and separated by SDS-PAGE. 32P-autoradiogram and Western blot analysis of 254-nm UV-crosslinked (+) and noncrosslinked (−) complexes for each indicated RBP candidate are shown. Known RBPs (Elav, Pum, Smg), served as positive controls. RNA-signals were compared against FLAG-IP of crosslinked parental S2 cells to estimate nonspecific signal. RBP candidates Wech, GlyP, Hsc70Cb, and CG6287 could not be verified. (C) Enrichment of uniquely aligned CLIP sequencing reads relative to matched total RNA input for Pep and CG3800. X-axis: fraction of reads per million (RPM). Y-axis: log2-transformed RPM ratio of CLIP vs input samples. (Circles, replicate 1; diamonds, replicate 2.) (D) 5-mer enrichment of randomly sampled aligned RBP CLIP reads relative to matched inputs. X-axis: frequency of 5-mer in 106 bases. Y-axis: 5-mer frequency ratio CLIP vs. input. (E) Enrichment analysis of aligned sequencing reads to mRNA subannotation categories. Sequencing reads were normalized to reads per kilo base per million (RPKM). Y-axis: log2-transformed ratio of annotation categories RPKMs in CLIP vs. input. (TES) transcription end site.











