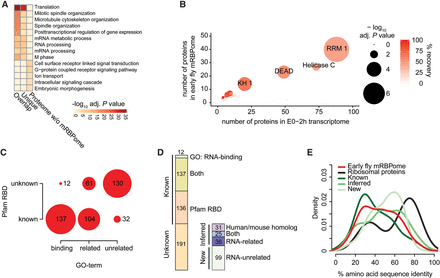
Characterization of the early fly mRBPome. (A) GO analysis showing the five most enriched gene ontology terms for molecular functions (GOMF) of the mRNA-bound proteins (overlap and unique) and the remaining proteins identified from whole embryos. P values were calculated by comparing against the early embryo transcriptome (0–2-h old embryos, all genes with FPKM > 0), adjusted for multiple testing with Benjamini-Hochberg and −log10-transformed. (B) Pfam protein domain enrichment of the early embryo mRBPome (n = 476) (y-axis) compared to the early embryo transcriptome (n = 7298) (x-axis). P values were calculated with Fisher's exact test and Bonferroni-corrected for multiple testing, indicated by circle size. Red shading indicates recovery percentages of expressed genes. (C) The intersection of proteins with known RNA-binding GO-term and Pfam RNA-binding domain (RBD) (list of RBDs and RNA-binding GO-terms described by Gerstberger et al. [2014]). (D) Proportions of proteins previously annotated as known RBPs by either GO-term or RBD (Known) and proteins not previously found to directly interact with RNA (Unknown). Unknown proteins contain homologs identified in mouse and human mRNA interactome studies and/or are part of the human RBP census, and proteins with RNA-related GO-terms (together referred to as ‘inferred RBPs’). ‘New’ refers to the 99 proteins undescribed in terms of RNA-binding. (E) Protein amino acid sequence identity to human homologous proteins using Ensembl Compara (Vilella et al. 2009). The early fly mRBPome without ribosomal proteins (n = 371) is depicted in subgroups containing: ribosomal proteins (n = 56), all known RBPs (n = 206), inferred RBPs (n = 80), and new RBPs (n = 85).











