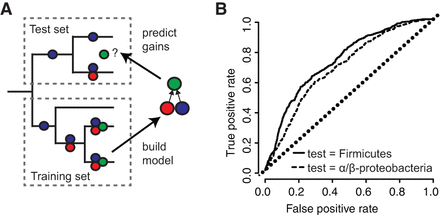
PGCE dependencies lead to taxonomically robust predictability of gene acquisition. (A) Workflow for predicting gene acquisition between clades of the tree. A training set is used to build a PGCE dependency model, which is then used to predict on which specific branches genes are likely to be gained (green circles), based on dependencies inferred from the training set (red and blue circles). (B) Performance of PGCEs in predicting gene acquisitions in two test sets (indicated clades of the prokaryotic tree). Areas under each curve: Firmicutes, 0.73; Alpha/Beta-proteobacteria, 0.68. The diagonal dotted line represents the performance of a purely random prediction. See also Supplemental Figure S9.











