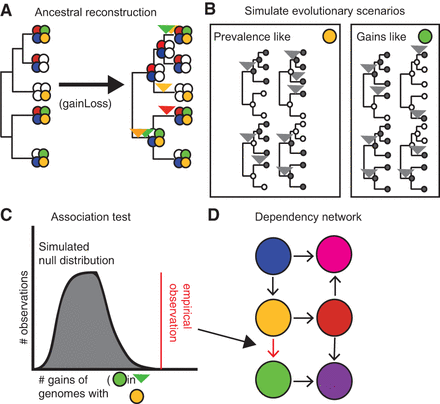
Workflow for deriving the PGCE network. (A) A model phylogeny and a set of gene presence/absence patterns at the tips are used to generate an ancestral reconstruction, from which gains are inferred. Filled circles represent the presence of a gene (distinguished by color), and empty circles represent absence of that gene. Inverted triangles represent points on the phylogeny where the gene of the indicated color is inferred to be gained. (B) Based on inferred gain and loss rates, many evolutionary scenarios are independently simulated and used as a null expectation for evolutionary independence. Filled circles indicate presence of the simulated gene, and empty circles indicate absence; inverted triangles represent gains of the simulated gene on the phylogeny. (C) A null distribution derived from simulated gene evolution is used to identify dependencies between real genes. (D) These dependencies are modeled as a network. Filled circles indicate genes (nodes); arrows indicate dependencies (edges).











