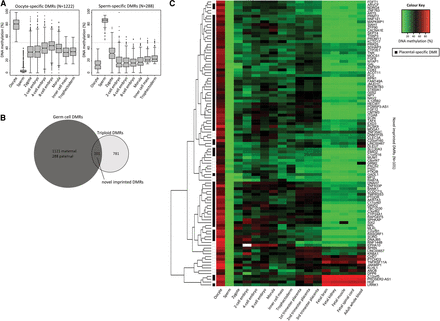
Identification of novel imprinted placental DMRs. (A) DNA methylation is shown for oocyte-specific (left panel) and sperm-specific (right panel) DMRs in human gametes and early embryonic stages (zygote, two-cell, four-cell, eight-cell, and morula stage embryos), ICM, and TE. DMRs were defined as CGIs that showed a >50% methylation difference between gametes, with intermediate methylation (15%–60%) in the ICM and TE. (B) The number of DMRs that overlap between those identified between human gametes and those identified between triploid placental samples is shown. (C) DNA methylation through early human development is shown for the 101 novel imprinted DMRs. DNA methylation for human gametes, early embryonic stages (zygote, two-cell, four-cell, eight-cell, and morula stage embryos), ICM, and TE is an average of CpG sites across each DMR, measured by RRBS. DNA methylation for placental villi, fetal tissues (brain, kidney, muscle, and spinal cord), and whole blood was an average of 450K array probes across each DMR. Seventy-two novel DMRs showed a placental-specific pattern of imprinting, defined as <25% or >75% methylation in somatic tissues and intermediately (25%–75%) methylated in placenta.











