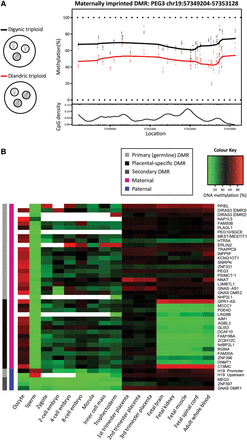
Characterization of known imprinted DMRs in human development. (A) Digynic triploid conceptions contain two maternal (white) and one paternal (gray) genome, while diandric triploid conceptions contain one maternal and two paternal genomes. By comparing digynic (N = 5) and diandric (N = 5) triploid placental villi, DMRs were identified, defined as ≥3 CpGs within 500 bp. An example plot shows a maternal DMR near PEG3, which is more highly methylated among digynic triploid samples (black line), compared to diandric triploid samples (red line) across 37 CpG sites over a ∼4-kb region. (B) DNA methylation through early human development is shown for 43 DMRs, identified between triploid samples that overlap previously reported imprinted DMRs (Court et al. 2014). DNA methylation for human germ cells, early embryonic stages (zygote, two-cell, four-cell, eight-cell, and morula stage embryos), inner cell mass, and trophectoderm is an average of CpG sites across each DMR measured by RRBS. DNA methylation for placental villi, fetal tissues (brain, kidney, muscle, and spinal cord), and whole blood was an average of 450K array probes across each DMR. Primary (germline) imprinted DMRs (light gray), placental-specific DMRs (black), and secondary DMRs (dark gray) are denoted. Parental origin of DNA methylation is designated as maternal (magenta) or paternal (blue). White boxes in the heat map indicate DMRs with no data.











