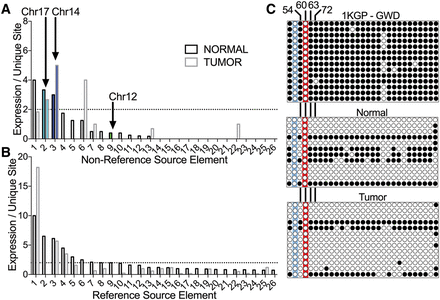
FL-L1Hs source element expression in normal and tumor tissues. The unique interior mutation profiles of FL-L1Hs elements in the patient's genome (Fig. 2) were used to quantify expression of the 31 nonreference FL-L1Hs source elements including the Chr 17, Chr 14, and Chr 12 source elements (A) and the remaining 264 reference FL-L1Hs source elements in the patient's genome, using strand-specific RNA-seq (B) (Supplemental Table S3; Methods). Expression for each element is depicted in the normal and tumor tissues as the mean number of independent reads covering all mutations unique to a FL-L1Hs source element. Expression of FL-L1Hs source elements that could not be differentiated from the expression of the surrounding gene in the same orientation were excluded from this analysis (A, n = 4; B, n = 37) (Supplemental Table S3). The horizontal dotted lines represent a cutoff where the mean = two traces per unique site, and a total of 10 elements were expressed above this level. (C) DNA methylation analysis of the Chr 17 source element promoter. The top panel displays bisulfite sequencing results for the control 1000 Genomes Project (1KGP) sample (GWD sample HG02583, which is heterozygous for the Chr 17 element; Coriell). The two panels below show the results of bisulfite sequencing in the normal and tumor tissues of the CRC patient. Each circle represents a CpG site in the promoter (for a total of 29 CpGs at positions: 21, 37, 54, 60, 63, 72, 102, 137, 155, 160, 164, 166, 171, 181, 205, 231, 251, 255, 269, 284, 293, 305, 317, 320, 327, 351, 363, 369, 377 relative to the reference L1.3 sequence; GenBank ID L19088). White circles indicate no DNA methylation; black circles indicate DNA methylation. The red highlighting indicates the CpG at position 60 that previously was shown to be critical for repression of the L1 promoter by DNA methylation (Hata and Sakaki 1997). This CpG is mutated in the Chr 17 element (G61A). The three remaining CpG sites that are critical for repression of the L1 promoter by DNA methylation also are indicated (positions 54, 63, 72) (Hata and Sakaki 1997). The blue highlighting indicates a CpG to TpG mutation at position 37 that destroys an additional CpG in the promoter region.











