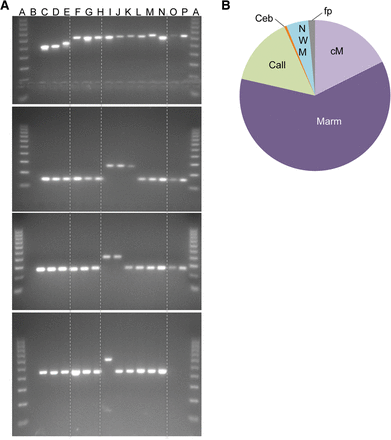
Phylogenetic distribution of Platy-1. (A) Four agarose gel chromatographs of our locus-specific phylogenetic analyses. An upper fragment indicates presence of a Platy-1 insertion; a lower fragment, absence. The vertical lines from left to right separate the outgroups from NWMs, Atelidae from Cebidae, and Cebidae from Pitheciidae. The gel chromatographs show (from left to right): (A) 100 bp ladder; (B) TLE; (C) human; (D) common chimpanzee; (E) African green monkey; (F) woolly monkey; (G) spider monkey; (H) red howler monkey; (I) common marmoset; (J) pygmy marmoset; (K) tamarin; (L) capuchin monkey; (M) squirrel monkey; (N) owl monkey; (O) titi; (P) saki. (For more detailed information regarding the species used, please see Supplemental Table S1A.) The top gel image shows a Platy-1 insertion shared across all NWMs. The gel chromatograph below shows an insertion specific to Callithrichinae, which is followed by a marmoset-specific insertion. The gel chromatograph on the bottom shows a common marmoset-specific Platy-1 insertion. (B) The phylogenetic results for our informative loci are shown in a pie chart: (Marm) marmosets; (cM) common marmoset; (fp) false positive; (NWM) New World monkey; (Call) Callithrichinae; (Ceb) Cebidae.











