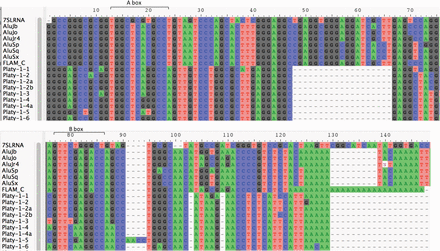
Alignment of Platy-1 with 7SL RNA and Alu elements. Shown is a multiple sequence alignment using MUSCLE (Edgar 2004) followed by manual curation of the oldest Platy-1 subfamilies with 7SL RNA, a selection of Alu consensus sequences, and FLAM. The alignment is visualized with AliView (Larsson 2014). Dashes indicate absence of the sequence. Also illustrated are the A box (consensus sequence: TRGYnnAnnnG) and B box (consensus sequence: GWTCRAnnCc). The tail of the Platy-1 sequence aligned equally well to the regions prior to the middle A-rich region and the 3′ end of an Alu element. In the latter case, the deletion may have been caused by recombination between homologous sequences. This alignment assumes that the element terminates at the middle A-rich region. An alternate alignment is provided in Supplemental Figure S4.











