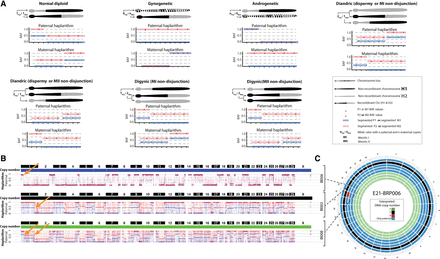
Haplarithmisis reveals abnormal fertilization (dispermy) and segregation of parental genomes in distinct cell lineages. (A) Schematic overview of haplarithm pattern for different genomic constitutions. The segmented M1, M2, P1, and P2 single-cell BAF values, as well as the distances between “M1 and M2” and “P1 and P2” values in maternal and paternal haplarithms, respectively, and the positioning of breakpoints, i.e., homologous recombinations (HR), represent inheritance of different parental homologous chromosomes in a single blastomere (for details, see Zamani Esteki et al. 2015). Note that for simplicity's sake, we show only one chromosome, and each pattern should be seen in all (or the majority) of chromosomes to be considered as a genome-wide event. (B) Genome-wide paternal haplarithms of three blastomeres derived from the same embryo (E21_BRP006) uncover different HR-sites in two cells (top and middle) and a combination thereof in the other cell (bottom). The orange arrows point to HR sites on Chr 1. Note that here, option 1 of siCHILD (see Methods) was applied as the paternal grandparents were used for phasing the paternal genome. (C) Circos plot illustrating the genome-wide interpreted copy-number profiles of all the single blastomeres derived from embryo E21_BRP006.











