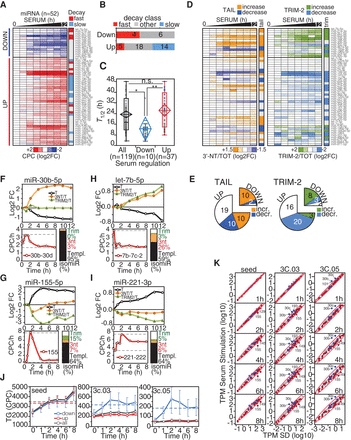
Regulation of miRNAs by decay dynamics following serum stimulation. (A) The heat-map shows fold-change (log2) of serum-regulated miRNAs (n = 52) over the 12-h time course following serum treatment. Data are the average of two independent biological replicas (shown in Supplemental Fig. S6A). “Fast” and “slow” decay miRNAs are indicated by red and blue, respectively. (B) The distribution of serum-regulated miRNAs among decay classes was analyzed by contingency (χ2 = 9.3, P = 0.0096). (C) Distribution of miRNAs half-lives (T1/2) within classes of serum-regulated miRNAs. The “ALL” class (N = 119) contains all the common miRNAs of the serum-regulation and decay data sets. (D) Heat-maps show the variation of tailing (3′-NT/TOT) and trimming (TRIM-2/TOT) frequencies over the 12-h time course, as in A. (E) Number of miRNAs with changes in tailing (χ2 = 12.0, P = 0.0025) and trimming (χ2 = 15.2, P = 0.0005) within classes of serum-regulated miRNAs. (F–I) Representative examples of “DOWN” (F,H) and “UP” (G,I) serum-regulated miRNAs are shown. For each miRNA, the regulation of templated (T), tailed (3′-NT/TOT), and trimmed (TRIM-2/TOT) forms is reported, along with the changes in the primary transcript (below; in red and measured as CPC/h). The bar plot summarizes the frequency of each miRNA isoform. (J) Mean target abundance (with SEM) of down-regulated (in blue), up-regulated (in red), or all miRNAs (in black) over the time course of serum stimulation. (K) Correlation between target:miRNA ratio (TPM) in growing versus serum-depleted fibroblasts. Straight lines with shaded confidence intervals highlight the linear correlation. Targets were further distinguished according to their complementarity to the miRNA 3′ end, as described in Figure 4, A and B.











