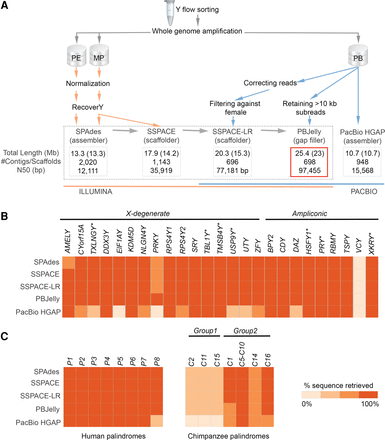
(A) The global workflow applied for the Y Chromosome assembly (see text for details). Four assemblies in the dotted frame are nested within each other. The best assembly is framed in red. (Orange) Illumina data; (blue) PacBio data. All assemblies were filtered against the reference female genome. The total (including Ns) and unambiguous (non-N, shown in parentheses) lengths are shown. N50 is the contig/scaffold length for which all contigs/scaffolds of that length or longer contain half of the assembly length. (B) Gene and (C) palindrome recovery. The heatmaps show how sequences homologous to 25 human genes, eight human palindromes, and 12 chimpanzee-specific palindromes were recovered in the assemblies (see Methods). Genes lost on the chimpanzee Y are marked with an asterisk.











