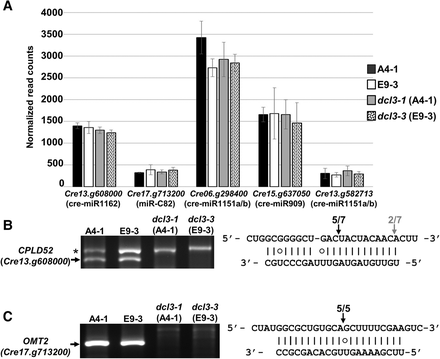
The effect of miRNA on mRNA accumulation. (A) Steady-state accumulation levels of previously reported miRNA targets (Molnár et al. 2007; Zhao et al. 2007) assessed as the number of normalized reads (y-axis) in RNA-seq data. Error bars for three independent samples are shown. The target genes with their corresponding miRNA are indicated. These miRNAs were predicted as either high confidence miRNAs (cre-miR1162, cre-miR1151a/b) or medium confidence miRNA (miR-C82) (see Table 1), with the exception of cre-miR909 that is a hairpin-derived siRNA also depleted in the dcl3 mutant background. (B) 5′RACE to test the specific cleavage of CPLD52 (Cre13.g608000) mediated by cre-miR1162. The asterisk indicates an unspecific PCR product. The PCR products were sequenced, and the right panel shows the 5′ terminus of these cleavage products aligned to the 5′ to 3′ mRNA sequence and the 3′ to 5′ miRNA. G:U base pairs are indicated by a circle. (C) 5′RACE to test the specific cleavage of OMT2 (Cre17.g713200) mediated by the miR-C82 (Zhao et al. 2007) with the 5′ terminus of these cleavage products aligned to the 5′ to 3′ mRNA sequence and the 3′ to 5′ miRNA as in B.











