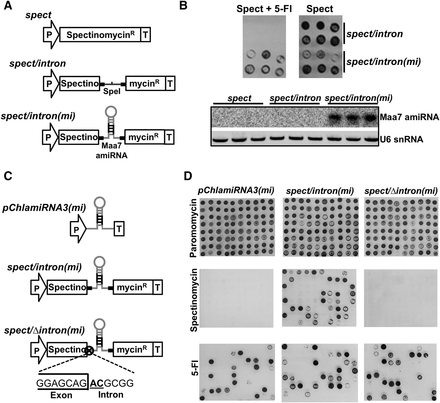
The cre-miR1157 is an intron-derived miRNA. (A) Schematic representation of constructs carrying the cre-miR1157 intron inserted into the spectinomycin resistance gene coding sequence. The cre-miR1157 intron was modified to either lack the miRNA stem–loop or carry an artificial miRNA against Maa7 in spect/intron and spect/intron(mi) plasmids, respectively: (P) Hybrid RBSC2/HSP70A promoter; (SpectinomycinR) recoded Escherichia coli-derived aadA coding gene; (T) RBSC2 transcription terminator; (SpeI) unique cleavage site for SpeI restriction enzyme; (Maa7 amiRNA) modified version of cre-miR1157 that carries a miRNA against Maa7. (B, top) Growth of the indicated transgenic lines in solid media carrying spectinomycin with/without 5-Fluorindole (5-Fl). (Bottom) Detection by Northern blot of the artificial miRNA against Maa7 in total RNA samples from the indicated lines (three independent lines per construct). (C) Schematic representation of constructs used to test the requirement of splicing for the expression of id-miRNA. The GT × AT point mutations in the exon/intron junction are indicated. These plasmids also carry the ParomomycinR cassette (equivalent to the cassette showed in Fig. 1A) to allow the primary selection of transgenic lines in paromomycin. (D) Growth of lines transformed with the indicated plasmids in solid media containing either paromomycin (test for plasmid integration), spectinomycin (test for splicing events), or 5-Fl (test for amiRNA production).











