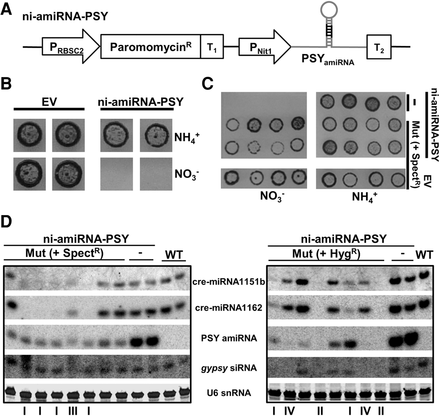
Screening and isolation of mutants affected in miRNA-mediated RNA silencing. (A) Schematic representation of the artificial miRNA construct used to transform the wild-type strain. Transgenic lines carrying this cassette were further screened by random insertional mutagenesis: (PRBSC2) RuBisCO small subunit (RBSC)2 promoter; (ParomomycinR) Streptomyces rimosus AphVIII coding gene; (T1) RBSC2 transcription terminator; (PNit1) nitrate reductase promoter; (PSYamiRNA) modified version of cre-miR1157 that carries a miRNA against the phytoene synthase; (T2) RLP12 transcription terminator. (B) Selective cell death of transgenic lines expressing the PSY amiRNA in the presence of nitrate, but not ammonium, as the sole nitrogen source. Transgenic lines carrying the empty amiRNA vector (EV) were used as control. (C) Growth in high light conditions of mutagenized (SpectR) and nonmutagenized (-) reporter lines (PSYamiRNA) in solid media containing either nitrate or ammonium as sole nitrogen source. Transgenic lines carrying the empty amiRNA vector (EV) and further transformed with the spectinomycin resistance cassette were used as control. (D) Detection by Northern blot of diverse small RNAs in total RNA samples from the indicated mutants and controls. These mutants were obtained by random insertional mutagenesis of either spectinomycin or hygromycin resistance cassettes. The mutants were grouped (I–IV) (Supplemental Table S1) based on the molecular phenotype. The two displayed mutants belonging to the group II correspond to the characterized mutant 47 (dcl3-2) and mutant 51 (dcl3-3).











