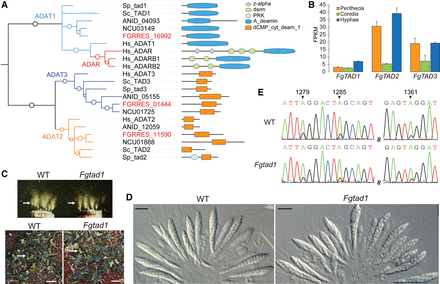
Evolution, expression, and function of ADATs in F. graminearum. (A) Phylogenetic tree of deaminase domain of fungal ADATs and ADARs constructed using PhyML3.1 (Guindon et al. 2010). The SH-like support of approximate likelihood ratios (aLRT-SH) is plotted as circles on the branches (only SH-like support >0.6 are shown). The prefixes for gene names or IDs are as follows: (ANID) Aspergillus nidulans; (FGRRES) Fusarium graminearum; (Hs) Homo sapiens; (NCU) Neurospora crassa; (Sc) Saccharomyces cerevisiae; (Sp) Schizosaccharomyces pombe. The domain structures are as follows: (A_deamin) adenosine-deaminase (PF02137); (dCMP_cyt_deam_1) cytidine and deoxycytidylate deaminase zinc-binding region (PF00383); (dsrm) double-stranded RNA binding motif (PF00035); (PRK) phosphoribulokinase (PF00485); (z-alpha) adenosine deaminase z-alpha domain (PF02295). (B) The expression level (fragments per kilobases of exons for per million mapped reads [FPKM]) of three ADAT genes of F. graminearum estimated with RNA-seq data of conidia, 24-h hyphae, and perithecia collected at 8 dpf. Error bars indicate standard deviations calculated from two biological replicates of RNA-seq data. (C) Mating cultures of the wild-type PH-1 (WT) and Fgtad1 deletion mutant were examined for ascospore release and cirrhus production. Arrows point to ascospore masses and cirrhi (ascospores oozing) ejected from perithecia. Bar, 1 mm. (D) Asci and ascospores formed by PH-1 and the Fgtad1 mutants were examined with 12-dpf perithecia. Bar, 20 μm. (E) Sequencing traces for the edited region of FgSSN3 (FGRRES_04484) amplified from RNA isolated from perithecia of PH-1 and Fgtad1 mutant. Black arrows mark the edited A's that have a mixed peak of A and G in sequencing traces.











