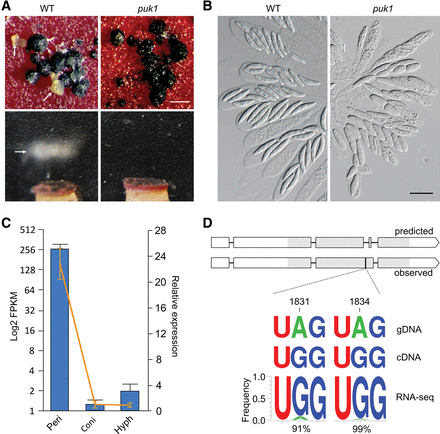
Function, expression, and RNA editing of PUK1. (A) Mating cultures of the wild-type PH-1 (WT) and puk1 mutant were examined for cirrhus production (upper) and ascospore release (lower). Arrows point to cirrhi (ascospores oozing) and ascospore masses ejected from perithecia. Bar, 1 mm. (B) Asci and ascospores formed by PH-1 and the puk1 mutant. Deletion of PUK1 affected ascospore morphology. Bar, 20 μm. (C) The expression level of PUK1 in conidia (Coni), 24-h hyphae (Hyph), and perithecia collected at 8 dpf (Peri). The bar chart represents the absolute expression level (log2 FPKM) in RNA-seq data, and the line is the relative expression level (2−ΔΔCt) assayed by qRT-PCR (the expression level of PUK1 in conidia arbitrarily set to one). Error bars indicate standard deviation calculated from two biological replicates of RNA-seq data or three biological replicates for qRT-PCR. (D) The gene structure and editing sites of PUK1. The gene model and coding region of PUK1 is different between automated annotation (predicted) and actual cDNA sequence (observed). Rectangle boxes are coding regions, and the protein kinase domain region is in gray. The corrected gene model contains two tandem stop codons, UA1831G UA1834G (1830–1835, marked with a black vertical line) in its coding region that is part of an intron introduced erroneously by automated annotation. A1831and A1834 in the genomic DNA (gDNA) were changed to G's in cDNA sequences by RNA editing. WebLogo shows the frequency of A-to-G variants at each site in RNA-seq reads.











