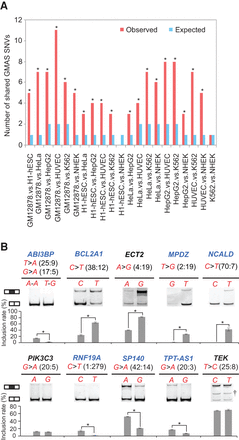
Splicing alteration by genetic variants is highly cell type independent. (A) The number of common GMAS SNVs between each pair of samples is shown (red bars). The expected number of shared GMAS SNVs for each pair of samples (blue bars) was calculated assuming an independent occurrence of each event in each sample. Enrichment of common GMAS SNVs between cell types was evaluated using hypergeometric test by comparing the observed and expected occurrences. (*) P < 0.05. (B) The same splicing assays as shown in Figure 2C were repeated by transfecting the minigenes (containing human iGMAS exons) into mouse 3T3 cells. The results are illustrated in the same way as in Figure 2C. Successfully validated cases are illustrated with the gene names in blue.











