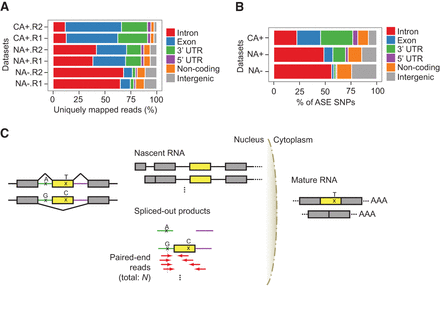
Compartment-specific RNA-seq enables coverage of intronic SNVs. (A) The genomic context of uniquely mapped reads of GM12878 CA+, NA+, and NA− RNA-seq data sets. Two biological replicates (R1 and R2) were analyzed. Noncoding refers to noncoding transcripts or genes. Reads that mapped to regions with multiple annotation categories were classified into one genomic context according to priorities given as follows: coding exon > 3′ UTR > 5′ UTR > intron > noncoding > intergenic. (B) Similar to A, the genomic context of SNVs with ASE patterns in GM12878 CA+, NA+, and NA− data sets. Biological replicates were combined. (C) The biological principle underlying iGMAS detection. In this hypothetical example, the yellow exon is alternatively spliced (AS) depending on the allele of the intronic SNV, with the A allele associated with exon inclusion and G allele associated with exon skipping. In the nucleus, NA− RNA-seq reads could originate from nascent RNA or spliced-out products. In the spliced-out products, the A and G alleles will reside, respectively, in the single intron and the intron–exon–intron molecule. In the nascent RNA, the A and G alleles are also present, which is not shown in the diagram. RNA-seq reads (red arrows) originating from spliced-out products covering the intronic SNV and neighboring exon or intron are enriched with the G allele, which can be analyzed to infer allele-specific regulation of splicing. An exonic SNV is also illustrated, whose T allele (the one in the same haplotype as the intronic A allele) is expected to be enriched in RNA-seq data of the cytoplasmic RNA (CA+). Note that exonic SNVs are not always present in iGMAS exons.











