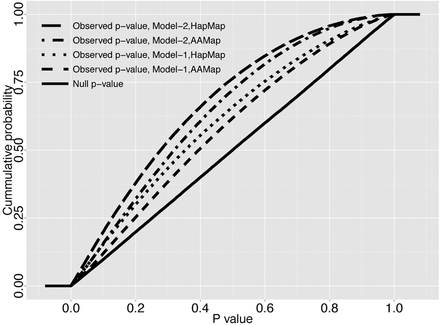
Significant excess of tests with very low P-values in the observed S* P-value distribution. Plotted is the observed (dashed lines) S* P-value distribution for the real data, calculated based on each of the four sets of whole-genome simulations. The four sets arise from the combination of the two demographic null models (Supplemental Fig. S2): Model-1 (the continuous asymmetric gene flow model) and Model-2 (the single-pulse admixture model); and the two genetic recombination maps: HapMap Yoruba map (HapMap) and African American map (AAMap). The solid line represents the null S* P-value distributions of the four whole-genome null simulation sets, derived by calculating P-values using a single randomly chosen simulation from each set. All four are uniformly distributed between 0 and 1 as expected. For the real data (dashed lines), all four analyses show a significant shift to small P-values in the observed S* P-value distribution (one-sided Mann-Whitney U test, P < 2.2 × 10−16), thus rejecting our demographic null hypotheses including no archaic admixture.











