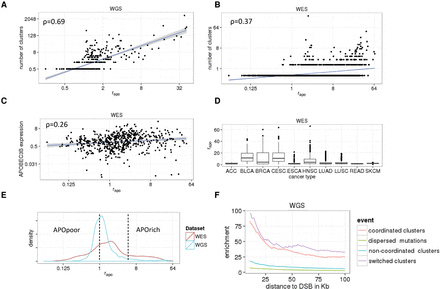
Properties of cancers associated with the APOBEC mutational signature (rapo). (A,B) Correlation between rapo and the number of kataegistic mutations +0.5 per tumor in WGS (A) and WES (B) data sets. (C) Correlation of rapo and the level of expression of APOBEC3B per tumor in the WES data set. (D) Box plots representing rapo per tumor type; WES data set: (ACC) adrenocortical carcinoma; (BLCA) bladder cancer; (BRCA) breast cancer; (CESC) cervical squamous cell carcinoma; (ESCA) esophageal carcinoma; (HNSC) head and neck squamous cell carcinoma; (LUAD) lung adenocarcinoma; (LUSC) lung squamous cell carcinoma; (READ) rectum adenocarcinoma; (SKCM) cutaneous melanoma. (E) Distributions of rapo in WGS and WES data sets. Tumors with rapo < 1 were categorized as APOpoor, and tumors with rapo > 5, as APOrich. (F) Enrichment of dispersed and clustered APOBEC signature mutations in the proximity of rearrangement breakpoints (used as a proxy for DSBs) in the WGS data set. The observed mutation rates in the proximity (up to 100 kb) of rearrangement breakpoints were compared with the average mutation rates per tumor in APOrich tumors.











