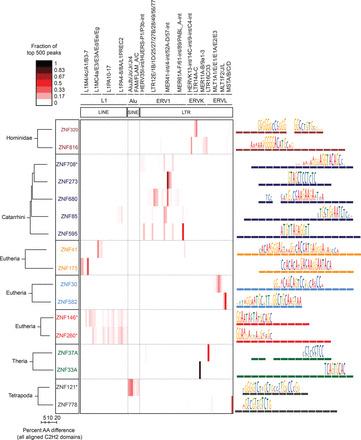
ERE binding pattern in seven groups of C2H2-ZF paralogs. The heat map (center) indicates the fraction of the top 500 ChIP-seq peaks overlapping each ERE (ERE classes indicated at top). Paralogs are grouped together in boxes (left) and their aligned C2H2-ZF domain structures are represented by colored rectangles (right) (Clustal Omega [v.1.2]) (Sievers et al. 2011). Asterisks indicate C2H2-ZF proteins that lack a KRAB domain. Taxon names at left indicate the most recent lineage where the paralogs share at least one homologous finger. Binding motifs (right) are positioned over the corresponding C2H2-ZF domains that recognize each triplet according to RCADE (Najafabadi et al. 2015a). Aligned C2H2-ZF domains of paralogs are displayed as dashed lines in the same color.











