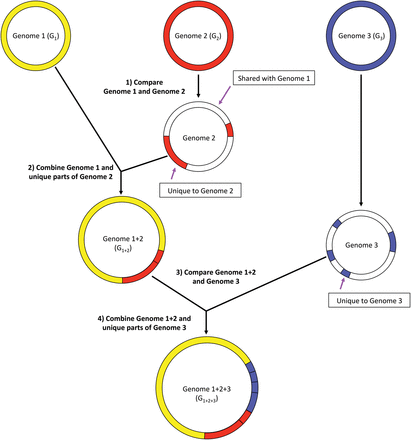
Compression of genome sequences before building the Centrifuge index. All genomes are compared and similarities are computed based on shared 53-mers. In the figure, genomes G1 and G2 are the most similar pair. Sequences of G2 that are ≥99% identical to G1 are discarded, and the remaining “unique” sequences from G2 are added to genome G1, creating a merged genome, G1+2. Similarity between all genomes is recomputed using the merged genomes. Sequences <99% identical in genome G3 are then added to the merged genome, creating genome G1+2+3. This process repeats for the entire Centrifuge database until each merged genome has no sequences ≥99% identical to any other genome.











