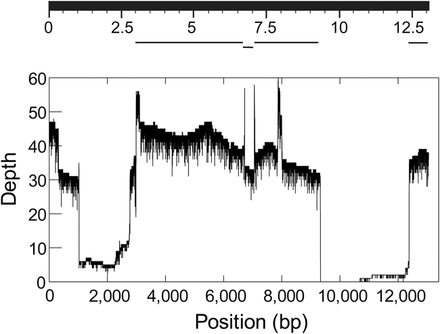
Sampling bias in the Drosophila PacBio data. (Top) BlastN search using the Y-linked kl-3 gene CDS as the query against a database of the MHAP-assembled Drosophila genome (Berlin et al. 2015). Note the large assembly gaps. (Bottom) Coverage depth of the same gene in the raw PacBio reads. Note that most of assembly gaps in the kl-3 gene actually were caused by low or absent coverage in the PacBio reads. The expected coverage depth is 45×. The Illumina coverage of the same region is fairly homogeneous (Supplemental Fig. S12).











