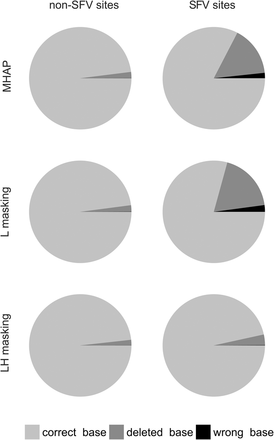
Read correction accuracy within a segmental duplication (human 10q11 region). Corrected reads were aligned with the original sequence, and each base of each read was scored as “correct” (light gray), “wrong” (black), or “deleted” (dark gray). “SFV sites” (for “sequence family variant”) are located within the segmental duplication, at positions where the two copies are different. “Non-SFV sites” are sites located within the segmental duplication and identical between the two copies or located outside the segmental duplication (they produce identical results and were lumped in the figure). Note that standard MHAP and MHAP with L-masking frequently fail at SFV sites, whereas LH-masking correctly handles them. Data from 450 sites of each type; reads were corrected with the default falconsense algorithm (for the pbdagcon correction, see Supplemental Fig. S10).











