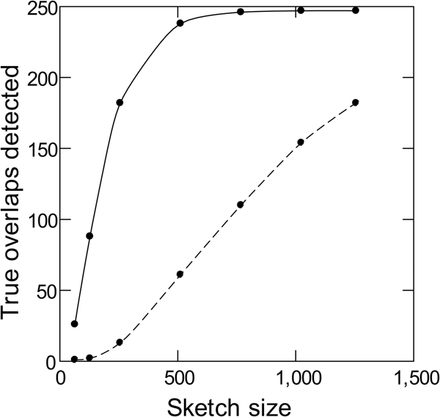
Figure 2.
Sensitivity of read overlap detection with and without k-mer validation. Simulated PacBio reads from E. coli (250 pairs of 10-kb sequences with 2-kb overlaps) were subjected to standard MHAP (dashed line) or MHAP with masking of low-frequency k-mers (solid line) for overlap detection. The reference list of valid k-mers came from Illumina reads.











