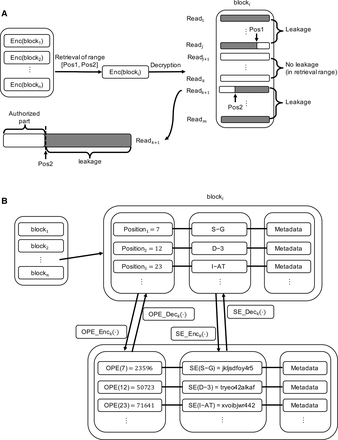
Solutions for encrypting genomic data. (A) A standard SE solution on CRAM-compressed data. It leads to potential data leakage when querying specific genomic regions. The encryption is performed over each individual data block. As a block in CRAM usually contains multiple sequencing reads, it is usually the case that the retrieved block will reveal reads or positions that are not in the query position range. (B) SECRAM encryption. Our solution encrypts each block in SECRAM format based on positions. Position information is encrypted with OPE; the compressed content at each position, with SE. This ensures that only information corresponding to the query position range is retrieved and decrypted. “OPE(POSITION)” represents the OPE ciphertext of POSITION, and “SE(VARIANT)” represents the SE ciphertext of VARIANT. Metadata are not encrypted (e.g., quality scores, mapping quality, read name). Note that OPE preserves the order of the positions, namely, 23596 < 50723 < 71641 because 7 < 12 < 23, but SE encrypts the original message to a random string, for example, from “S-G” to “jkljsdfoy4r5.”











