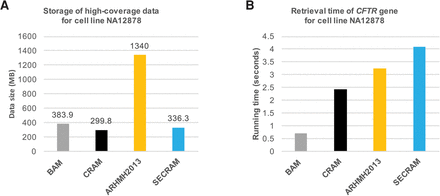
(A) Storage and (B) runtime on high-coverage clinical data. The data come from a public cell line (NA12878), based on a gene panel that includes the CFTR gene, queried for diagnosing cystic fibrosis. The average coverage of these data is 1035×, containing more than 4 million reads of length around 300 bases. ARHMH2013 (Ayday et al. 2013) is a privacy-preserving solution on BAM files that does not address the compression requirement. We observe a consistent compression performance of SECRAM on high-coverage clinical sequence data. Moreover, querying for the CFTR gene on SECRAM takes less than twice the time on the nonencrypted CRAM.











