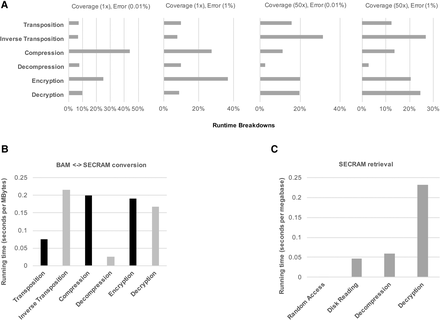
Runtime analysis of SECRAM system for storing and retrieving genomic data. (A) Runtime breakdowns on simulated data sets for the six most important procedures in SECRAM: transposition, inverse transposition, compression, decompression, encryption, and decryption, relative to total runtime (=100%). The experiments were repeated for four simulated data sets with low (1×)/high (50×) coverages and low (0.01%)/high (1%) error rates. In all cases, the runtime cost of encryption/decryption is comparable with other necessary procedures, showing that enforcing security does not result in significant performance overhead. (B) Conversion time on real data sets (same data sets as those in Fig. 2B). It shows the average conversion time between the BAM and SECRAM formats on real data sets, running with a single thread on a machine equipped with Mac OS X Yosemite system and 3.1-GHz Intel Core i7 processor. The black bars (transposition, compression, encryption) represent the three steps of conversion from BAM to SECRAM, whereas the light gray bars (decryption, decompression, inverse transposition) represent the three steps of conversion from SECRAM to BAM. Each step takes <0.25 sec per megabyte of data. (C) Retrieval time on real data sets (same data sets as those in Fig. 2B). It shows the average runtime cost of retrieving data within a range of 1 million genomic positions. The actual data size corresponding to 1 million positions depends on the coverage. Shown are experiments on real data sets from Figure 2B with a coverage of about 3× and a size of slightly >1 megabyte.











