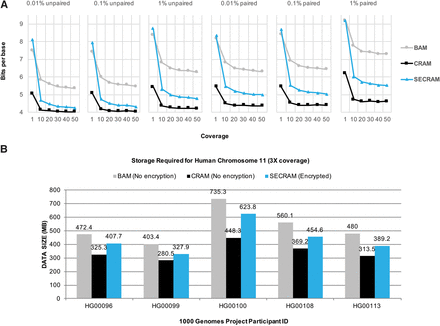
Storage analysis of compression algorithms for genomic data. (A) Storage comparison on simulated data sets. It shows the average number of bits per base (compression ratio) for three storage formats: BAM, CRAM, and SECRAM. The results are based on simulated data sets with different coverages (from one to 50) and error rates (0.01%, 0.1%, 1%) for both paired and unpaired data. (B) Storage comparison on Chromosome 11 from the 1000 Genomes Project participants with an average coverage of 3×. Only SECRAM is an encrypted format, whereas both BAM and CRAM are in plaintext. Both A and B show that the compression ratio of SECRAM is between that of BAM and that of CRAM.











