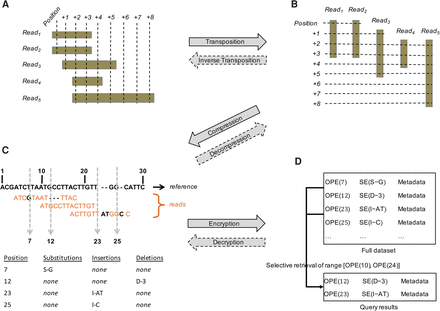
SECRAM framework for compressed, encrypted storage of genomic data. (A) The sequencing read-based format used in BAM is transposed into a genome position-based format (B). The read-based format can be reconstructed from the position-based format via reverse transposition. (C) The position-based storage is compressed and decompressed using a reference-based compression technique. In the table, “S-G” stands for substitution with base “G”, “D-3” stands for deletion of three bases, and “I-AT” stands for insertion of two bases “AT.” (D) The compressed position-based storage is encrypted to generate the final SECRAM format using order-preserving encryption (OPE) and traditional symmetric encryption (SE) scheme. “OPE(POSITION)” represents the OPE ciphertext of POSITION, and “SE(VARIANT)” represents the SE ciphertext of VARIANT. Metadata are not encrypted (e.g., quality scores, mapping quality, read name). The compressed format is recovered from the encrypted format by running the respective decryption algorithms. Our encryption enables efficient selective retrieval. For instance, if a user wants to retrieve data in the range [10, 24], the database executes a normal search between OPE(10) and OPE(24) based on the order-preserving property of OPE, and in the shown example, two positions, OPE(12) and OPE(23), are returned.











