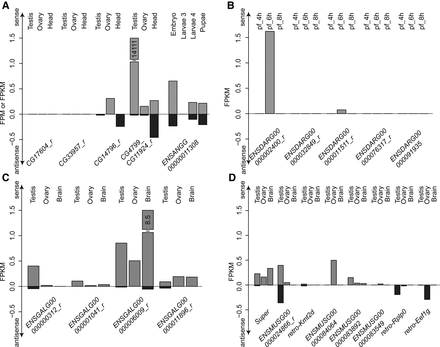
Strand-specific RNA-seq-based quantification. On the basis of newly generated and public strand-specific RNA-seq data, we quantified the expression of LTR-mediated retrocopies in fruit fly and mosquito (A), zebrafish (B), chicken (C), and mouse (D). For fruit fly, given the high similarity between polymorphic retrocopies and parental genes, only switch points spanning reads were counted, and FPM was calculated. For all other cases, FPKM was calculated. For mosquito, embryo, the third instar larvae, the fourth instar larvae, and pupae were profiled. For zebrafish, 4-, 6- and 8-h post fertilization (pf) stages were profiled.











