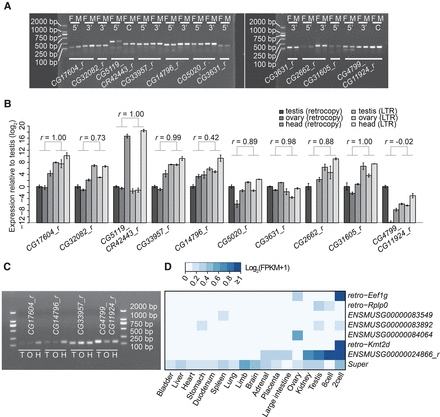
The expression of retrocopies in fruit fly and mouse. (A) Whole-body RT-PCR results for 10 retrocopies validated in Supplemental Figure 2, which are expressed in both male (M) and female (F). The primer sequences and expected product sizes are listed in Supplemental Table 9. For the chimeric genes, CG5119_CR42443_r and CG4799_CG11924_r, we also designed primers flanking the fusion point between the two parental genes, the product of which is marked with “C.” (B) Tissue profiling via qRT-PCR. CG2662_r has no detectable expression in ovary. The Pearson correlation (r) between the retrocopies and corresponding retrotransposons across the three tissues is displayed above. (C) Tissue level RT-PCR results for four of 10 retrocopies encoded by line 208 across Testis (T), Ovary (O), and Head (H). (D) Expression profile of retrocopies encoded by the mouse genome. Tissues are hierarchically clustered on the basis of expression similarity across genes. Retrocopies are not clustered but instead are sorted by age as approximated by nucleotide divergence. To reveal relative moderate expression, the color-code scheme is truncated at the FPKM cutoff of 1 (i.e., all expression higher than 1 is shown in dark blue).











