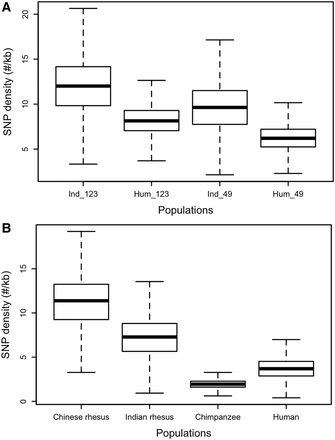
(A) Density of autosomal SNVs observed in rhesus (Indian rhesus only, 123 samples) and human (1000 Genomes Project data, phase 1, 123 randomly selected samples). Ind_123 indicates results for 123 rhesus samples; Hum_123 for 123 human samples; Ind_49 indicates 49 low-coverage (average 9.47×) Indian rhesus samples; Hum_49 for 49 human samples with the average coverage 9.44×. (B) Density of autosomal SNVs observed in Chinese rhesus (sample size n = 9), Indian rhesus (nine samples randomly sampled from 123 Indian rhesus samples), chimpanzee (nine samples randomly sampled from 10 samples; downloaded from http://panmap.uchicago.edu), and human (1000 Genomes Project data, phase 1, nine samples randomly sampled from 1092 samples).











