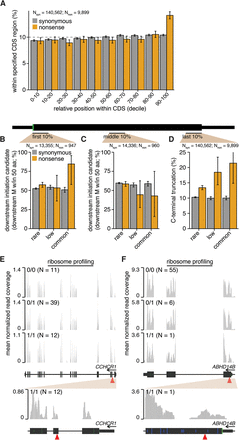
N- and C-terminal protein truncation enable translation of nonsense variant–containing mRNAs. (A) Positional distribution of variants within their host transcripts. Plot indicates the percentages of variants lying within the indicated deciles of their host CDSs’ lengths. Error bars, 95% confidence intervals as estimated by the binomial proportion test. Percentages of variants lying within the first 10% (B) or middle 10% (C) of their host CDS's length that have a downstream methionine (M) within 50 amino acids. Plot restricted to variants that lie within genes containing nonsense variants. Error bars, 95% confidence intervals as estimated by the binomial proportion test. (D) Percentages of variants lying within the last 10% of their host CDS's length. Plot restricted to variants that lie within genes containing nonsense variants. Ribosome profiling read coverage of nonsense variants lying within CCHCR1 (E) and ABHD14B (F). Units, data normalization, and notation are as in Figure 3. (Blue bars) CTG or GTG noncanonical start codons (no downstream ATG codons are present within the plot region). (Full/half-height blue bars) CTG or GTG codons that are/are not within a Kozak consensus context, defined as RnnSTGG (R = A/G, S = C/G).











