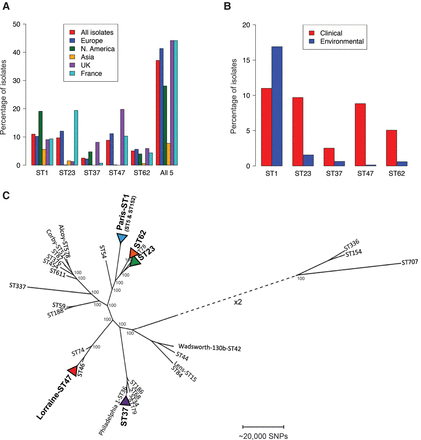
Geographical and environmental distribution of the five major disease-associated STs and their phylogenetic context within the species. (A) The percentage of clinical isolates submitted to the ESGLI sequence-based typing (SBT) database for L. pneumophila that are sequence types (STs) 1, 23, 37, 47, or 62, or from all five STs combined, are shown. Data are shown from different regions and countries where the numbers of isolates submitted were deemed sufficient for comparison. Data are based on a total of 6116 epidemiologically unrelated clinical isolates (i.e., including only one isolate from clusters/outbreaks) submitted to the database, 4785 of which were detected in Europe, including 541 in the United Kingdom and 2313 in France, as well as 801 in North America and 323 in Asia. (B) The percentage of clinical and environmental isolates from each of the five STs from the total number of clinical and environmental isolates submitted to the SBT database. Data are based on a total of 6116 epidemiologically unrelated clinical and 2826 unrelated environmental isolates (i.e., including only one isolate from clusters/outbreaks) submitted to the database. (C) Maximum likelihood phylogeny, generated by mapping sequence reads to the Corby reference genome, of representatives from each of the five dominant disease-associated STs together with an additional 27 isolates that are representative of the species diversity. The size of the alignment was 3,576,470 bp. Generally, the five STs are found in separate major clades of the species except for ST23 and ST62 that share a closer phylogenetic relationship. The scale represents the number of SNPs. Bootstrap values, derived from 1000 resamples, are shown for the major nodes of the tree.











