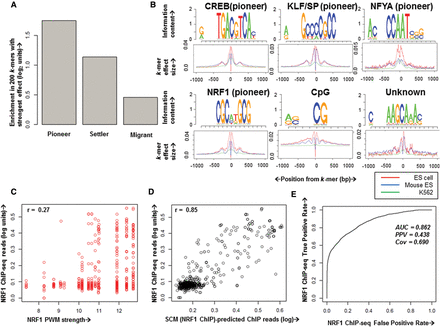
Chromatin accessibility arises from synergistic interactions, largely among pioneer TFs. (A) Enrichment of the 200 k-mers with strongest mESC SCM effect sizes in similarity to pioneer, settler, and migrant TF motifs. (B) Example position weight matrix (PWM) motifs derived from clustering the 500 k-mers with strongest mESC SCM effect size. Below the PWM are merged spatial k-mer effect sizes for all k-mers contributing to the motif within ±1000 bp of the k-mer in hESC (red), mESC (blue), and K562 (green), showing the common effects of k-mers in these cell types. Names above correspond to high-confidence database matches with TF motifs when known, and known pioneer TFs are denoted. (C,D) NRF1 ChIP-seq reads from 1-kb regions surrounding above-threshold NRF1 motif matches on held-out Chromosome 18 (y-axis) plotted versus NRF1 PWM strength (C, x-axis) or SCM (NRF1 ChIP)-predicted ChIP reads in the region (D, x-axis). Pearson's correlation coefficients are shown above each plot. (E) Receiver–operator curve (ROC) showing predictive accuracy of a SCM trained on NRF1 ChIP-seq data at predicting held-out NRF1 ChIP-seq peaks.











