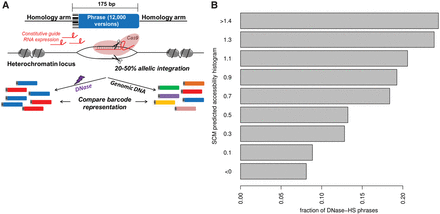
SCMs predict sequence-dependent chromatin accessibility in a high-throughput SLOT screen. (A) In SLOT, a library of 12,000 175-bp DNA sequences containing 100-bp variable phrases is PCR amplified to add a 67-bp homology arm to each end. mESCs stably expressing a guide RNA targeting a natively heterochromatin region are co-electroporated with the DNA library, Cas9, and additional guide RNA, resulting in a population of cells in which 20%–50% of alleles at a single genomic locus have phrase incorporation. Comparison of phrase representation between genomic DNA and DNase I hypersensitive DNA reveals the subset of phrases encoding chromatin accessibility in this controlled genomic context. (B) Fraction of phrases, grouped by their overall SCM-predicted chromatin accessibility (y-axis), that are DNase I hypersensitive in a SLOT assay (x-axis). The units shown are only appropriate for trend comparison.











