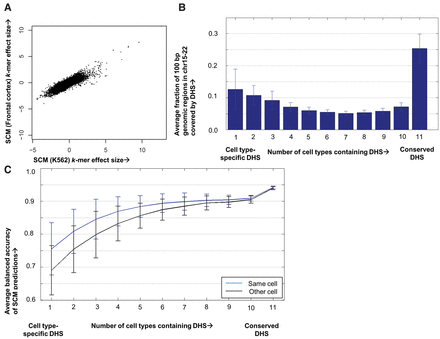
SCMs are more accurate at predicting DNase I hypersensitive regions that are active in multiple cell types. (A) Summed k-mer effect sizes for each k-mer in SCMs trained on K562 (x-axis) versus human frontal cortex (y-axis). (B) Histogram showing the fraction of the genomic space covered by DNase I hypersensitive sites (DHS) in a single cell type that are hypersensitive in 10 other human cell types. (C) Histogram showing the average balanced accuracy of SCM DHS predictions binned according to the cell-type specificity of DHS activity.











