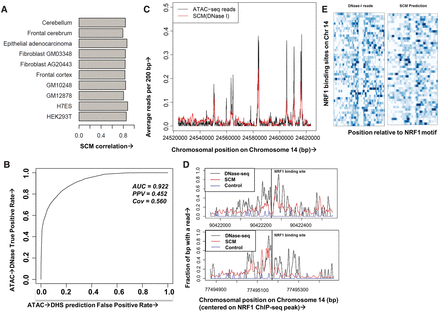
SCMs predict chromatin accessibility at base pair resolution across cell types and data types. (A) Pearson correlation coefficients on held-out Chromosome 14 DNase-seq data for SCMs trained on DNase-seq from 10 cell types. (B) Receiver–operator curve (ROC) showing predictive accuracy of a SCM trained on GM12878 DNase-seq data at predicting held-out GM12878 ATAC-seq peaks. (C) Example human GM12878 held-out genomic region showing ATAC-seq reads (black) and reads predicted from a DNase-seq trained SCM (red), both smoothed at 200 bp. (D) Example human GM12878 held-out genomic region showing 10-bp smoothed DNase-seq reads (black), SCM-predicted reads (red), and reads from a control model trained on IMR-90 naked DNA DNase-seq data (blue) surrounding two NRF1 binding sites (vertical black lines denote binding call). (E) Heatmap showing clear footprints for both DNase-seq and SCM at NRF1 binding sites on Chromosome 14.











