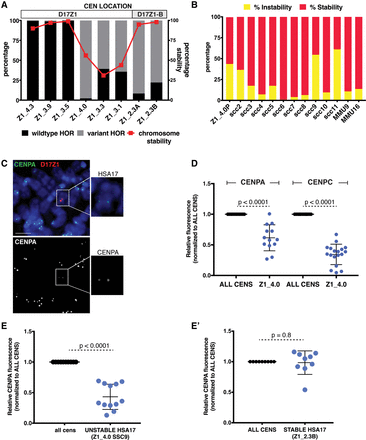
Centromeres assembled at variant D17Z1 arrays are less stable than homogeneous wild-type arrays. (A) The proportion of variation (wild-type versus variant HORs) in a subset of HSA17s is plotted with stability of the HSA17 (red line). For each line, chromosome stability was determined using FISH with D17Z1 probes and counting the number of HSA17s in 200 cells (stability was defined by maintenance of HSA17 ploidy in each line). When the centromere formed on a large, homogeneous array of D17Z1, such as in Z1_4.3, Z1_3.9, and Z1_3.5, the HSA17 was extremely stable in mitosis. Similarly, when the centromere assembled at D17Z1-B in lines Z1_2.3A and Z1_2.3B (highly variant D17Z1 arrays), HSA17 was very stable. However, when the centromere was assembled on D17Z1 arrays that had moderate or extreme variation, HSA17 was mitotically unstable. Centromere location (D17Z1—white, D17Z1-B—gray) on each HSA17 is denoted above the plot. (B) Line Z1_4.0 had the most variant (98%), yet active, D17Z1 array in our data set, and this HSA17 exhibited chromosome instability. The parental Z1_4.0 (Z1_4.0P) was subcloned to produce multiple, independent versions of the HSA17 (single-cell clones, SCC); subcloning could also account for aberrant behavior in a single cell line that did not reflect inherent behavior of the HSA17. The single-cell clones showed varying levels of chromosome instability, indicating that the unstable phenotype was inherent to this HSA17. The stability of two mouse chromosomes (MMU9, MMU16) was measured to account for genetic background effects that might alter the stability of all chromosomes. (C) CENPA and CENPC (Supplemental Fig. S4E) immunostaining (green) was combined with FISH using D17Z1 probe p17H8 (red) to quantitate the amount of centromere proteins on unstable Z1_4.0. Insets show the HSA17 alone and a single channel image of CENPA staining on the HSA17. Scale bar, 15 µm. (D) The amount of CENPA and CENPC on unstable Z1_4.0 was plotted compared to all other centromeres in the cell. Fluorescence from all centromeres (ALL CENS) was normalized to one, and the fluorescence at Z1_4.0 was calculated according to this normalized value. The amount of CENPA at the Z1_4.0 centromere was half of the amount at all other centromeres in the cell; CENPC was reduced by more than 50%. (E) The unstable Z1_4.0 single cell clone SSC9 also showed reduced amounts of CENPA. By comparison, the amount of CENPA on Z1_2.3B, a stable HSA17 that has a variant D17Z1 array but assembles the centromere at D17Z1-B, was comparable to all the other centromeres in the cell (E′).











